Pde1a
Mus musculus phosphodiesterase 1A, calmodulin-dependent (Pde1a)
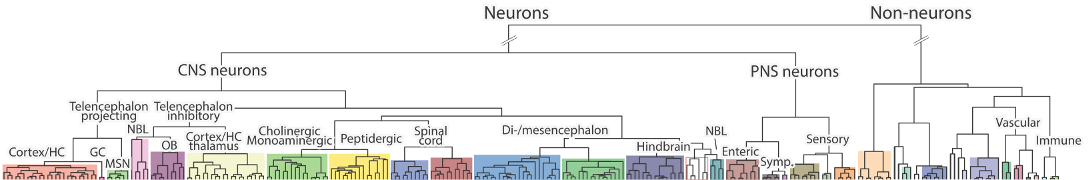

Clusters expressing Pde1a
Table shows clusters that express the gene at a trinarization score >= 0.95
| Expression | Index | Symbol | Description | Region |
| 1.6982 | 2 | TEGLU3 | Excitatory neurons, cerebral cortex | Telencephalon |
| 2.8577 | 3 | TEGLU2 | Excitatory neurons, cerebral cortex | Telencephalon |
| 3.4545 | 4 | TEGLU20 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.1059 | 5 | TEGLU11 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.3238 | 6 | TEGLU12 | Excitatory neurons, cerebral cortex | Telencephalon |
| 4.1282 | 7 | TEGLU10 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.6089 | 8 | TEGLU9 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.3523 | 9 | TEGLU8 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.7051 | 10 | TEGLU7 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.5984 | 12 | TEGLU13 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.9714 | 13 | TEGLU14 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.4286 | 14 | TEGLU5 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.1148 | 15 | TEGLU16 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.8941 | 17 | TEGLU17 | Excitatory neurons, cerebral cortex | Telencephalon |
| 0.5434 | 18 | TEGLU18 | Excitatory neurons, cerebral cortex | Telencephalon |
| 2.1573 | 19 | TEGLU19 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.9509 | 20 | TEGLU22 | Excitatory neurons, amygdala | Telencephalon |
| 2.875 | 21 | TEGLU21 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 2.1777 | 22 | TEGLU4 | Excitatory neurons, cerebral cortex | Telencephalon |
| 1.1978 | 23 | TEGLU24 | Excitatory neurons, hippocampus CA1 | Telencephalon |
| 2.2169 | 24 | TEGLU23 | Excitatory neurons, hippocampus CA3 | Telencephalon |
| 0.3264 | 30 | MSN4 | D1 medium spiny neurons, striatum | Telencephalon |
| 0.5923 | 51 | TEINH19 | Hippocamposeptal projection, cortex/hippocampus | Telencephalon |
| 2.1688 | 52 | TEINH21 | Sleep-active, long-range projection interneurons, cortex/hippocampus | Telencephalon |
| 0.6906 | 56 | TEINH20 | Inhibitory interneurons, hippocampus | Telencephalon |
| 0.4212 | 61 | TEINH11 | R-LM border Cck interneurons, cortex/hippocampus | Telencephalon |
| 2.438 | 67 | TECHO | Cholinergic interneurons, telencephalon | Telencephalon |
| 3.8333 | 68 | DECHO1 | Cholinergic neurons, septal nucleus, Meissnert and diagonal band | Diencephalon |
| 1.0962 | 69 | HBCHO4 | Afferent nuclei of cranial nerves III-V | Rhombencephalon |
| 0.4861 | 70 | HBCHO3 | Afferent nuclei of cranial nerves VI-XII | Rhombencephalon |
| 1.2308 | 73 | HYPEP7 | Pmch neurons, hypothalamus | Diencephalon |
| 0.617 | 75 | MEGLU14 | Glutamatergic projection neurons of the raphe nucleus | Mesencephalon |
| 0.4762 | 77 | MBDOP2 | Dopaminergic neurons, ventral midbrain (SNc, VTA) | Mesencephalon |
| 1.6 | 78 | HBSER1 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 1.2477 | 79 | HBSER2 | Serotonergic neurons, hindbrain | Rhombencephalon |
| 0.3636 | 105 | SCINH3 | Inhibitory neurons, spinal cord | Spinal cord |
| 0.47 | 120 | HBGLU2 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.8164 | 135 | DECHO2 | Cholinergic neurons, habenula | Diencephalon |
| 1.4615 | 136 | HBGLU1 | Excitatory neurons, hindbrain | Rhombencephalon |
| 0.6649 | 153 | TEINH1 | Inhibitory neurons, pallidum | Telencephalon |
| 0.7727 | 160 | HBINH2 | Inhibitory neurons, hindbrain | Rhombencephalon |
| 0.5 | 161 | HBCHO1 | Cholinergic neurons, hindbrain | Rhombencephalon |
| 0.7835 | 189 | ENT8 | Cholinergic enteric neurons, VGLUT2 | Neural crest |
| 0.629 | 192 | SYNOR2 | Noradrenergic neurons, sympathetic | Neural crest |
| 1.2133 | 196 | SYCHO2 | Cholinergic neurons, sympathetic | Neural crest |
| 0.5983 | 203 | PSPEP4 | Peptidergic (PEP1.1), DRG | Neural crest |
| 0.449 | 250 | ENMFB | Enteric mesothelial fibroblasts | Neural crest |
| 0.432 | 252 | VLMC2 | Vascular leptomeningeal cells | Neural crest |
| 0.2711 | 256 | VSMCA | Vascular smooth muscle cells, arterial | Mesoderm |
Pde1a is a marker of the following clusters
| Expression | Index | Symbol | Description | Region |
| 4.1282 | 7 | TEGLU10 | Excitatory neurons, cerebral cortex | Telencephalon |